Study population
This study is a part of a Virtual Consortium for Translational/Transdisciplinary Environmental Research (ViCTER) funded by the National Institute of Environmental Health Sciences. ViCTER consortium’s project (R01 ES032176), using a longitudinal cohort (NCT03281850), where participants were recruited between 2010 and 2014 from central Arkansas. Eligibility criteria were maternal age over 21 years, either actively planning or already pregnant within the first 10 weeks of gestation, second parity singleton pregnancies, and BMI between 18.5 and 35 kg/m2. Exclusion criteria comprised preexisting medical conditions, sexually transmitted diseases (STDs), medication influencing fetal growth, substance use, and pregnancy complications. Gestational diabetes was an exclusion criterion in our study and therefore was not present in the analytic sample. Placentas with severe pathologies were excluded. The final cohort enrolled 300 pregnant women, with placental samples obtained from 152 participants at term within 30 min of delivery.
Placental collection
All placental samples were collected within 30 min and processed within 2 h of delivery. After removing the umbilical cord and fetal membranes, samples of placental tissues were collected from the chorionic villous core following removal of the chorionic plate. The tissue underlying the chorionic plate consists of chorionic villous tissue and is of fetal origin. Samples were collected at 6 random sites (~ 1 sq. in) and washed thrice to remove maternal blood. To ensure comprehensive representation of the highly heterogenous placenta, ~1 g tissue from these 6 random sites was pooled, pulverized in liquid nitrogen and flash frozen. Samples were stored at -80 °C until DNA isolation.
DNA methylation analysis and processing
Genomic DNA extraction was done with PureLink DNA isolation reagents (Thermo Fisher). DNA was quantified with Qubit dsDNA quantification assays (Thermo Fisher, Waltham, MA, USA), then aliquoted into standardized concentrations to allow for a total mass of 500 ng of DNA. Samples were randomly distributed across 96-well plates, rows, and chips to reduce the potential for batch effects. Bisulfite conversion was done with the EZ DNA Methylation Kit (Zymo Research, Irvine, CA) and CpG-specific DNAm was quantified using the Illumina MethylationEPIC Beadarray (Illumina, San Diego, CA) at the Emory Integrated Genomics Core. To reduce the technical variation and probe-type bias, functional normalization and beta mixture quantile normalization were used19,20. All samples passed quality control steps. There was no evidence of batch effect via principal component analysis.
Gestational age acceleration
The epigenetic gestational age of 152 samples was estimated utilizing the R package planet15. This package comprises three distinct placental epigenetic clocks: the robust placenta clock (RPC), control placental clock (CPC), and refined RPC15. We focused on CPC due to its training on placental samples designated as controls, making it more representative of our study population compared to RPC, which was trained using methylation profiles from placentas with varied pregnancy conditions15. To generate GAA values, we regressed epigenetic GA on reported GA via linear regression, then extracted the residuals.
Body measurements
Weight, height, fat mass, and lean mass gain were assessed at 11 postnatal time points (2 and 4 weeks, 2, 3, 4, 5, 6, 9, 12, 18 and 24 months) using standardized methodologies reported previously21. Child body composition was evaluated at 11 postnatal timepoints, using quantitative nuclear magnetic resonance (QMR, EchoMRI-AH small, Echo Medical System, Houston, TX), which was previously validated in this population21.
Statistical analyses
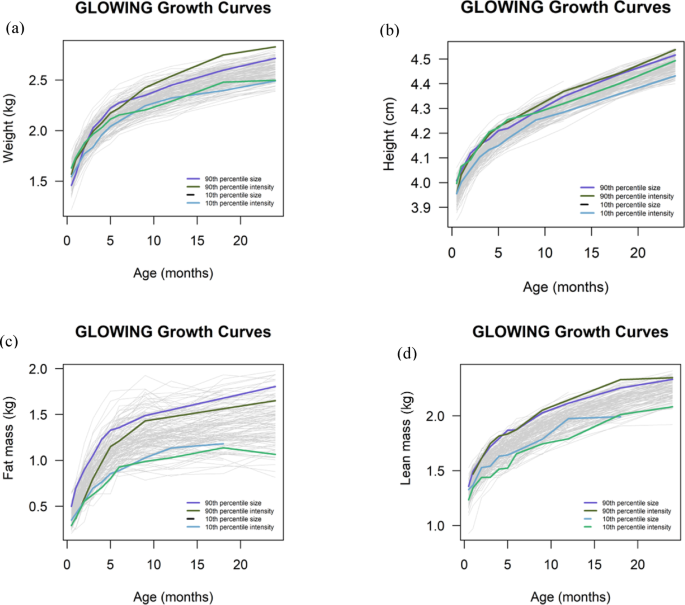
All statistical analyses were conducted using R (version 4.2.2). We utilized the SuperImposition by Translation and Rotation (SITAR) model to characterize postnatal trajectories of weight, height, fat mass and lean mass from 2 weeks through 24 months of age. The SITAR model functions as a shape invariant framework that yields a mean curve alongside sets of three parameters per individual (size, tempo, and velocity), summarizing the individual growth trajectories22. Individual curves are aligned with the mean curve (across all participants) through vertical shifts (representing variations in mean size) and horizontal shifts (reflecting differences in age at peak growth velocity, or tempo). Additionally, adjustments (termed intensity) to the temporal scale capture individual variations in growth velocity. These three parameters are estimated as random effects for each study participant. The tempo parameter was not able to be generated from our data due to missing values, leaving intensity and size parameters for analysis. To date, SITAR has been commonly used to understand the growth trajectory of weight and height23,24, however, this study also used the same approach to demonstrate and understand the trajectories of fat mass and lean mass in children.
Size and intensity values for weight, height, fat mass and lean mass were regressed on GAA (independent variable) while adjusting for potential confounding variables that could affect the associations such as maternal age, maternal education, gestational age, gestational weight gain, maternal race, and placental cell type proportions. A statistical significance of 0.05 level was used for all the results.
link





